![[Nanoprobes Tech Support (14k)]](../Images/logo_tech.gif)
Updated: March 6, 2008
PRODUCT INFORMATION
EnzMet for General Research Applications
[EnzMet for General Research Applications product page]

| Product Name: |
EnzMet for General Research Applications |
| Catalog Number: |
6010 |
| Revision:: |
1.1 (January 2008) |
Technical Assistance Online
![[2080-PDF]](../Images/pdf.gif) Instructions (PDF) Instructions (PDF)
The kit of EnzMet for General Research Applications is designed for use with horseradish peroxidase (HRP) for the sensitive and direct visualization of antigens, proteins, and other targets in general research applications. EnzMet produces black, sharply defined and non-diffusing stains with high sensitivity and resolution. Staining is permanent, and does not fade with time.
The following materials are sufficient for 150 tissue sections, based upon 200 µL per section:
| EnzMet General Detect A | 18 mL |
| EnzMet General HRP Detect B | 6 mL |
| EnzMet General HRP Detect C | 6 mL |
Refrigerate at 4°C. The product is shipped at ambient temperature.
EnzMet detection utilizes probes labeled with peroxidase enzymes to reduce silver ions to elemental silver, resulting in the deposition or accumulation of silver metal particles at sites labeled by peroxidases.1,2 The silver deposits can be directly visualized by eye or microscopy, or detected by other methods for the detection of target proteins,3 DNA4,5 or RNA sequences and other specific binding targets in matrix or biological specimens. EnzMet has also been used for the conductimetric detection of oligonucleotides on conductive array biochips,6 and the dense, highly localized silver deposits also provide intense, high-resolution labeling by electron microscopy.7
EnzMet generates dense, sharply defined, non-diffusing black signals with low background. Its high sensitivity and signal-to-noise ratio enables the detection of targets at low concentration. The non-diffusing and sharply defined stains provide superior spatial resolution for differentiation of targets. The black silver signals are readily distinguishable, and are not soluble in either aqueous or organic solution.
|
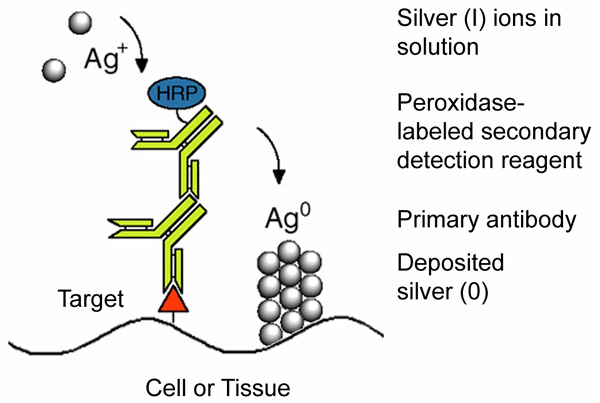
Figure 1:
Mechanism of the enzyme metallographic process showing enzyme-catalyzed deposition of metal from solution at a target site labeled with a primary antibody and peroxidase-labeled secondary antibody.
|
Contents
All EnzMet reagents and other required materials should be equilibrated to room temperature prior to use. All incubations with EnzMet detection should be performed at room temperature with gentle agitation. EnzMet solution B may turn become slightly yellow-colored during storage. However, the color change will not affect its performance.
For HRP detection, follow your standard procedure until incubation with the HRP-conjugated antibody or HRP reagent. After the incubation with the HRP-conjugated antibody or probe, follow these steps:
- Wash with buffer containing 0.1% Tween 20 for 3 x 5 minutes.
Note: Phosphate buffered saline, tris buffered saline or other wash buffers can be used. Including 0.1 % (w/v) Tween-20 in the wash buffer was found to be helpful in reducing non-specific binding.
- Wash with deionized water for 3 x 5 minutes.
- Tap off excess water. Incubate your sample with 1 volume of EnzMet Detect A. Incubate for 4 minutes.
Note: Excess water can lead to the dilution of EnzMet reagents, resulting in weak staining and results which are difficult to reproduce.
- Add 1 volume of EnzMet Detect B to Detect A, and gently mix Solutions A and B. Incubate for 4 minutes.
- Add 1 volume of EnzMet Detect C to the mixture of A and B, and gently mix Solutions A, B and C. Incubate for 6 - 15 minutes, or until satisfactory staining is achieved.
Note: The EnzMet incubation time mainly depends on the target concentrations and staining temperature. Longer incubation may be needed for visualizing low concentration targets. However, longer incubation may lead to some non-specific background staining. The variation of EnzMet staining temperature can affect its silver deposition rate. Lower temperature slows down the deposition process, and thus a longer staining time may be required to reach a certain degree of staining density and sensitivity.
- Wash with deionized water for 3 x 5 minutes.
- Air-dry your sample.
Contents
EnzMet detection produces black, sharply defined and non-diffusing stains at the sites of antigens or other targets.
For immuno- HRP detection, we recommend blocking with 5% nonfat dried milk prior to incubation with antibodies. The concentrations and incubation times for primary and secondary antibodies should be tested and adjusted to find the best signal-to-noise ratio. Including 1% nonfat dried milk in incubations with primary and secondary antibodies may be helpful in reducing non-specific background staining.
Contents
- 1. Hainfeld; J. F.; Eisen; R. N.; Tubbs; R. R.; and Powell; R. D.: Enzymatic Metallography: A Simple New Staining Method. Microsc. Microanal.; 8 (Suppl. 2: Proceedings); Lyman; C. E.; Albrecht; R. M.; Carter; C. B.; Dravid; V. P.; Herman; B.; and Schatten; H. (Eds.); Cambridge University Press; New York; NY, 916 CD (2002).
Full paper (HTML)
- Hainfeld, J. F.; Powell, R. D., and Hacker, G. W.: Nanoparticle Molecular Labels. In: Nanobiotechnology, Mirkin, C. A., and Niemeyer, C. M. (Eds): Wiley-VCH, Weinheim, Germany; ch. 23, pp. 353-386 (2004).
- Tubbs, R.; Pettay, J.; Powell, R.; Hicks, D. G.; Roche, P.; Powell, W.; Grogan, T, and Hainfeld, J. F.: High-Resolution Immunophenotyping of Subcellular Compartments in Tissue Microarrays by Enzyme Metallography. Appl. Immunohistochem. Mol. Morphol., 13, 371-375 (2005).
- Tubbs, R.; Pettay, J.; Hicks, D.; Skacel, M.; Powell, R.; Grogan, T., and Hainfeld, J.: Novel bright field molecular morphology methods for detection of HER2 gene amplification. J. Mol. Histol., 35, 589-594 (2004).
- Powell, R. D.; Pettay, J. D.; Powell, W. C.; Roche, P. C.; Grogan, T. M.; Hainfeld, J. F., and Tubbs, R. R.: Metallographic in situ hybridization. Hum. Pathol., 38, 1145-1159 (2007).
- Moller, R.; Powell, R. D.; Hainfeld, J. F., and Fritzsche, W.: Enzymatic control of metal deposition as key step for a lowbackground electrical detection for DNA chips. Nano Lett., 5, 1475-1482 (2005).
- Powell, R.; Joshi, V.; Thelian, A.; Liu, W.; Takvorian, P.; Cali, A., and Hainfeld, J.: Light and Electron Microscopy of Microsporida using Enzyme Metallography. Microsc. Microanal., 12 (Suppl. 2: Proceedings); Kotula, P.; Marko, M.; Scott, J.-H.; Gauvin, R.; Beniac, D.; Lucas, G.; McKernan, S., and Shields, J. (Eds.), Cambridge University Press, New York, NY, 424CD (2006)
Full paper (HTML)
Technical Assistance Available.
For a complete list of references citing this product, please visit our References page on our web site.
Contents
|